ACER scientists are looking at the role that genetic diversity played in determining the response of coastal plant and animal species to the oil spill. How do they know individual plants or animals are genetically different? We have chatted previously about gene sequencing, but another technique they can use is SNP analysis.
SNP stands for single nucleotide polymorphism. Recall that a nucleotide is a chemical base in a DNA (or RNA) molecule. For DNA, 4 nucleotides occur, designated by the first letter of their name (A for adenine, G for guanine, C for cytosine, T for thymine). Poly is a prefix meaning many and the root morph means form. Thus, single nucleotide polymorphism refers to the variation in which base, G, C, T, or A occurs at a single location in a strand of DNA. In doing SNP analysis, scientists are looking for the degree of variation in nucleotides in a specific section of DNA.

What difference would result from having a different single nucleotide at a particular location on a DNA molecule? The answer is – it depends. A DNA molecule is a long strand of nucleotide bases (coiled into a 3d shape in a living cell). The strand is divided into sections called genes; genes direct the synthesis of specific proteins. Regions of DNA containing genes are called coding regions. In between the genes are sections of DNA that are not involved with protein synthesis. These regions are referred to as non-coding regions. Nucleotides in non-coding regions may have no function or they may control other aspects of the gene to protein process, such as how fast (or how slow) it occurs, what other factors must be present for it to occur, or even how fast the message ‘make this protein’ lasts in a cell. Thus, some SNPs have no impact while others affect protein synthesis and may determine how the cell (and organism) functions. For example, it is a SNP in a coding region that is associated with the disease cystic fibrosis and a SNP in a non-coding region that is associated with how susceptible individuals are to the hepatitis C virus.
Returning to coastal habitats, their plant and animal species and ACER research, a SNP analysis tells us how genetically diverse a group of individuals of the same species is. Recall that marsh plants typically grow clonally; individual plants growing from the same rhizome will be genetically identical. Picking 2 marsh plants for an experiment that are growing from the same rhizome would ensure genetic similarity. However, since most animals reproduce sexually (mixing genetic materials from 2 genetically different individuals), scientists need to have another way to measure genetic differences – a SNP analysis.
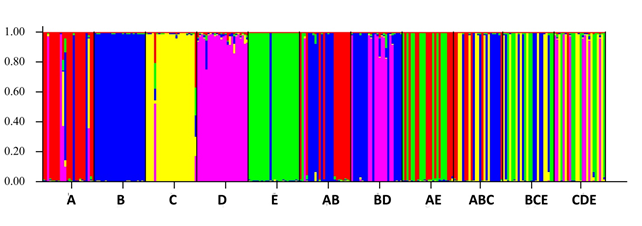
ACER Post-doctoral research associate Meagan Schrandt used a SNP analysis to assess genetic diversity among oysters in her experiments investigating their response to oil or oil and dispersant mixture. In collaboration with the Auburn University Shellfish Laboratory, she isolated several genetically different groups of oyster spat and then conducted a series of experiments measuring oyster growth and survivorship while varying oyster genetic diversity, salinity and oil exposure. Does this sound like a lot of work? You bet it was: the experiments required 330 individual tanks of water and 16,500 juvenile oysters! While her data analysis is still underway, she did share a portion of the results from her SNP analysis. In the figure below, letters represent groups of oysters and colors represent genetic diversity. Thus, there were 11 groups of genetically different oysters and within each group, genetic diversity was distinct from that of the other groups. Preliminary results show that survivorship varied with level of diversity, and higher survivorship was observed in tanks with more genetically diverse oyster populations.